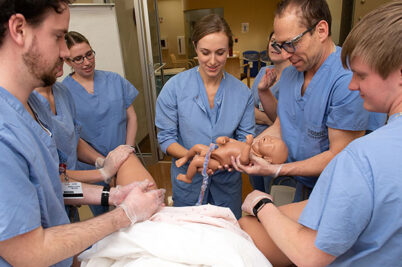
Training doctors for rural practice helps meet statewide need
Dr. Michelle Clark-Forsting always knew she wanted to be a doctor. Growing up in Alma Center, Wisconsin, she worked during the school year as a nursing assistant at nearby Black River Falls Memorial Hospital. Physicians there were her role models. In 2005, she applied to medical school at the University of Wisconsin.